Complete Chloroplast Genomes of Tulipa turkestanica and T. biflora (Liliaceae): Structural Conservation and Repeat Variation in Central Asian TulipsПолные геномы хлоропластов Tulipa turkestanica и T. biflora (Liliaceae): структурная консервативность и вариативность повторяющихся последовательностей в тюльпанах Центральной АзииTulipa turkestanica және T. biflora (Liliaceae) хлоропласттарының толық геномдары: Орталық Азия қызғалдақтарындағы құрылымдық сақталу және қайталанатын вариация.
Abstract
The genus Tulipa is taxonomically complex and highly diverse in Central Asia, yet genomic resources for many wild species remain limited. We sequenced and assembled the complete chloroplast genomes of Tulipa turkestanica and T. biflora, two morphologically variable species from Uzbekistan, to investigate plastome structure, repeat composition, and phylogenetic relationships. Both genomes exhibited conserved quadripartite organization with nearly identical sizes (152,022–152,025 bp) and each encoding 84 protein-coding genes, reflecting high structural stability within the genus. Comparative analyses revealed strong A/T bias in simple sequence repeats (SSRs) in both species. However, T. turkestanica displayed a higher number and greater diversity of SSR motifs, including species-specific dinucleotide repeats (particularly AT and AG/CT motifs), suggesting differential microsatellite accumulation during evolutionary divergence. Phylogenomic reconstruction based on complete chloroplast genomes provided robust resolution within subgenus Eriostemones, placing T. biflora with T. sogdiana and T. turkestanica with T. buhseana, both with strong bootstrap support (BS = 100). This topology contrasts with previous nuclear marker-based phylogenies, indicating potential influence of introgression and incomplete lineage sorting, a pattern commonly observed in Liliaceae. The petD-rpoA intergenic region exhibited the highest nucleotide diversity (π = 0.01625), representing a promising marker for population genetic studies. While plastome structure is highly conserved in Tulipa, repeat variation provides informative markers for species differentiation and evolutionary studies. These genomic resources enhance our understanding of tulip systematics and provide valuable tools for conservation genetics in these threatened wild species.
Род Tulipa является таксономически сложным и очень разнообразным в Центральной Азии, однако геномные ресурсы для многих диких видов остаются ограниченными. Мы секвенировали и собрали полные геномы хлоропластов Tulipa turkestanica и T. biflora, двух морфологически изменчивых видов из Узбекистана, чтобы исследовать структуру пластома, состав повторяющихся последовательностей и филогенетические отношения. Оба генома продемонстрировали консервативную четырехчастную организацию с почти идентичными размерами (152 022–152 025 п.н.) и каждый из них кодировал 84 белковых гена, что отражает высокую структурную стабильность внутри рода. Сравнительный анализ выявил сильный перекос A/T в простых повторяющихся последовательностях (SSR) у обоих видов. Однако у T. turkestanica было обнаружено большее количество и большее разнообразие мотивов SSR, включая видоспецифичные динуклеотидные повторы (особенно мотивы AT и AG/CT), что свидетельствует о дифференцированном накоплении микросателлитов в ходе эволюционной дивергенции. Филогенетическая реконструкция на основе полных геномов хлоропластов обеспечила надежное разрешение в подроде Eriostemones, поместив T. biflora с T. sogdiana и T. turkestanica с T. buhseana, оба с сильной поддержкой бутстрепа (BS = 100). Эта топология контрастирует с предыдущими филогенетическими деревьями, основанными на ядерных маркерах, что указывает на потенциальное влияние интрогрессии и неполного сортирования линий, что часто наблюдается в семействе Liliaceae. Межгенная область petD-rpoA продемонстрировала наибольшее нуклеотидное разнообразие (π = 0,01625), что делает ее многообещающим маркером для популяционно-генетических исследований. Хотя структура пластома в Tulipa высоко консервативна, вариации повторений предоставляют информативные маркеры для дифференциации видов и эволюционных исследований. Эти геномные ресурсы улучшают наше понимание систематики тюльпанов и предоставляют ценные инструменты для генетики сохранения этих угрожаемых диких видов.
Tulipa тұқымдасы Орталық Азияда таксономиялық тұрғыдан күрделі және өте алуан түрлі, бірақ көптеген жабайы түрлер үшін геномдық ресурстар шектеулі болып қала береді. Біз Өзбекстаннан шыққан морфологиялық тұрғыдан өзгермелі екі түр - Tulipa turkestanica және T. biflora - хлоропласт геномдарының толық геномдарын секвенирлеп, құрастырдық, бұл пластом құрылымын, қайталану құрамын және филогенетикалық байланыстарды зерттеуге мүмкіндік берді. Екі геном да бірдей өлшемдегі (152,022–152,025 б.п.) төрт бөлікті ұйымдасуды көрсетті және әрқайсысы 84 ақуызды кодтайтын генді кодтайды, бұл тұқым ішіндегі жоғары құрылымдық тұрақтылықты көрсетеді. Салыстырмалы талдаулар екі түрде де қарапайым тізбек қайталануларында (SSR) күшті A/T ауытқуын анықтады. Дегенмен, T. turkestanica эволюциялық дивергенция кезінде микросателлиттің дифференциалды жинақталуын қоса алғанда, SSR мотивтерінің саны мен әртүрлілігін көбірек көрсетті. Толық хлоропласт геномдарына негізделген филогеномдық қайта құру Eriostemones кіші тұқымдасында сенімді ажыратымдылықты қамтамасыз етті, T. biflora-ды T. sogdiana-мен және T. turkestanica-ды T. buhseana-мен орналастырды, екеуі де күшті bootstrap қолдауымен (BS = 100). Бұл топология бұрынғы ядролық маркерге негізделген филогениялармен қарама-қайшы келеді, бұл интрогрессияның және толық емес тектік сұрыптаудың ықтимал әсерін көрсетеді, бұл Liliaceae-де жиі байқалатын үлгі. petD-rpoA интергендік аймағы ең жоғары нуклеотидтік әртүрлілікті көрсетті (π = 0.01625), бұл популяциялық генетикалық зерттеулер үшін перспективалы маркер болып табылады. Пластом құрылымы Tulipa-да жоғары деңгейде сақталғанымен, қайталанатын вариация түрлердің дифференциациясы мен эволюциялық зерттеулер үшін ақпараттық маркерлерді қамтамасыз етеді. Бұл геномдық ресурстар қызғалдақ систематикасын түсінуімізді жақсартады және жойылып бара жатқан жабайы түрлерде сақтау генетикасы үшін құнды құралдарды ұсынады.
IntroductionВведениеКіріспе
Tulipa L. (Liliaceae) is a species-rich and taxonomically complex genus comprising approximately 97 described species worldwide (POWO, 2025). Mountainous Central Asia represents the primary center of origin and diversification of the genus and is recognized as a global biodiversity hotspot (Myers et al., 2000; Botschantzeva, 1962). Early floristic surveys documented 63 wild Tulipa species in this region (Vvedensky and Kovalevskaya, 1971), while subsequent taxonomic revisions have substantially expanded this number (Zonneveld, 2009; de Groot and Zonneveld, 2024; Asatulloev et al., 2023).
Persistent taxonomic uncertainty in Tulipa arises from pronounced morphological variation, frequent interspecific hybridization, and historical classification inconsistencies (Hall, 1940; Christenhusz et al., 2013; Dekhkonov et al., 2022). The complex geological history of Central Asia, including mountain uplift and climatic oscillations, has further promoted species diversification and endemism (Miao et al., 2012), supporting the hypothesis that the region represents the evolutionary cradle of tulips (Botschantzeva, 1962).
Despite their ecological and ornamental value, many Tulipa species are experiencing population declines, with 62 species currently listed in the IUCN Red List (2022). Among them, Tulipa biflora and T. turkestanica are widespread yet taxonomically challenging species characterized by high morphological variability and unresolved phylogenetic relationships. The scarcity of genomic resources has limited detailed evolutionary and comparative investigations for these taxa.
Chloroplast genomes provide powerful tools for resolving phylogenetic relationships and conducting comparative genomic analyses in plants. Their conserved structure, moderate evolutionary rate, and predominantly uniparental inheritance make them particularly valuable for systematic and evolutionary studies. Here, we present and compare the complete chloroplast genomes of T. biflora and T. turkestanica to elucidate their genomic features, clarify evolutionary relationships, and advance the systematics of Tulipa in Central Asia.
Tulipa L. (Liliaceae) — это богатый видами и таксономически сложный род, включающий приблизительно 97 описанных видов по всему миру (POWO, 2025). Горная Центральная Азия является основным центром происхождения и диверсификации рода и признана глобальной горячей точкой биоразнообразия (Myers et al., 2000; Botschantzeva, 1962). Ранние флористические исследования задокументировали 63 диких вида тулипы в этом регионе (Введенский и Ковалевская, 1971), в то время как последующие таксономические пересмотры существенно расширили это число (Зонневельд, 2009; де Гроот и Зонневельд, 2024; Асатуллоев и др., 2023). Сохраняющаяся таксономическая неопределенность в отношении тулипы возникает из-за выраженной морфологической изменчивости, частой межвидовой гибридизации и исторических несоответствий в классификации (Холл, 1940; Кристенхус и др., 2013; Дехконов и др., 2022). Сложная геологическая история Центральной Азии, включая поднятие гор и климатические колебания, способствовала дальнейшему разнообразию видов и эндемизму (Miao et al., 2012), что подтверждает гипотезу о том, что этот регион является эволюционной колыбелью тюльпанов (Botschantzeva, 1962).
Несмотря на их экологическую и декоративную ценность, многие виды тюльпанов испытывают сокращение численности, и в настоящее время 62 вида занесены в Красный список МСОП (2022). Среди них тюльпаны двуцветковые (Tulipa biflora) и турецкие тюльпаны (T. turkestanica) являются широко распространенными, но таксономически сложными видами, характеризующимися высокой морфологической изменчивостью и неразрешенными филогенетическими связями. Нехватка геномных ресурсов ограничивает детальные эволюционные и сравнительные исследования этих таксонов.
Геномы хлоропластов предоставляют мощные инструменты для разрешения филогенетических связей и проведения сравнительного геномного анализа растений. Сохранившаяся структура, умеренная скорость эволюции и преимущественно однородительское наследование делают их особенно ценными для систематических и эволюционных исследований. В данной работе мы представляем и сравниваем полные хлоропластные геномы T. biflora и T. turkestanica, чтобы прояснить их геномные особенности, уточнить эволюционные взаимосвязи и продвинуть систематику Tulipa в Центральной Азии.
Tulipa L. (Liliaceae) - бүкіл әлемде сипатталған шамамен 97 түрді қамтитын түрге бай және таксономиялық тұрғыдан күрделі тұқымдас (POWO, 2025). Таулы Орталық Азия тұқымдастың шығу тегі мен әртараптануының негізгі орталығы болып табылады және жаһандық биоәртүрліліктің ыстық нүктесі ретінде танылған (Myers et al., 2000; Botschantzeva, 1962). Ертедегі флористикалық зерттеулер осы аймақта 63 жабайы қызғалдақ түрін тіркеді (Введенский және Ковалевская, 1971), ал кейінгі таксономиялық түзетулер бұл санды айтарлықтай кеңейтті (Зонневелд, 2009; де Грут және Зонневелд, 2024; Асатуллоев және т.б., 2023).
Қызғалдақ-дағы тұрақты таксономиялық белгісіздік айқын морфологиялық вариациядан, жиі түраралық будандастырудан және тарихи жіктеу сәйкессіздіктерінен туындайды (Холл, 1940; Кристенхус және т.б., 2013; Дехконов және т.б., 2022). Орталық Азияның күрделі геологиялық тарихы, соның ішінде тау көтерілуі және климаттық ауытқулар, түрлердің әртараптануы мен эндемизмін одан әрі дамытты (Miao et al., 2012), бұл аймақ қызғалдақтардың эволюциялық бесігі деген болжамды растайды (Botschantzeva, 1962).
Экологиялық және сәндік құндылығына қарамастан, көптеген Tulipa түрлері популяцияның азаюын бастан кешіруде, қазіргі уақытта 62 түрі IUCN Қызыл тізіміне енгізілген (2022). Олардың ішінде Tulipa biflora және T. turkestanica кең таралған, бірақ таксономиялық тұрғыдан күрделі түрлер болып табылады, олар жоғары морфологиялық өзгергіштікпен және шешілмеген филогенетикалық байланыстармен сипатталады. Геномдық ресурстардың тапшылығы бұл таксондар үшін егжей-тегжейлі эволюциялық және салыстырмалы зерттеулерді шектеді.
Хлоропласт геномдары филогенетикалық байланыстарды шешу және өсімдіктерде салыстырмалы геномдық талдаулар жүргізу үшін қуатты құралдарды ұсынады. Олардың сақталған құрылымы, орташа эволюциялық жылдамдығы және негізінен бір аталық мұрагерлігі оларды жүйелі және эволюциялық зерттеулер үшін ерекше құнды етеді. Мұнда біз T. biflora және T. turkestanica хлоропласттарының толық геномдарын көрсетіп, салыстырамыз, олардың геномдық ерекшеліктерін анықтаймыз, эволюциялық байланыстарын нақтылаймыз және Орталық Азиядағы Tulipa-ның систематикасын жетілдіреміз.
Materials and MethodsМатериалы и методыМатериалдар мен әдістер
Plant Material and DNA Sequencing
Fresh leaf material of T. biflora and T. turkestanica was collected from natural populations in Uzbekistan. Voucher specimens were deposited in the TASH herbarium, and leaves were preserved in silica gel prior to DNA extraction. Genomic DNA was isolated using the DP305 Plant Genomic DNA Kit (Tiangen, China) following the manufacturer's protocol.
Sequencing libraries were prepared using the NEBNext Ultra™ DNA Library Prep Kit for Illumina (NEB, USA). DNA was fragmented to approximately 350 bp, followed by end repair, adapter ligation, PCR amplification, and purification using AMPure XP beads. Library quality was assessed using the Agilent 5400 Fragment Analyzer, and quantification was performed by qPCR. Paired-end sequencing (2 × 150 bp) was conducted on Illumina platforms at Novogene (Beijing, China).
Genome Assembly and Annotation
Clean reads were assembled into complete chloroplast genomes using the GetOrganelle pipeline v1.7.5 (Jin et al., 2020). Chloroplast genome annotation was performed in Geneious Prime v2023.1.2 using Tulipa buhseana (GenBank accession NC_052014) as a reference. All gene annotations were manually curated to verify gene boundaries and correct potential errors. Chloroplast genome maps were generated using OGDRAW v1.1 (Lohse et al., 2007).
Comparative and Phylogenetic Analyses
Simple sequence repeats (SSRs) in the chloroplast genomes were identified using the MISA web tool (Beier et al., 2017), with thresholds set to detect mono-, di-, tri-, tetra-, penta-, and hexa-nucleotide motifs with minimum repeat units of 10, 5, 4, 3, 3, and 3, respectively. Nucleotide diversity (π) across complete chloroplast genomes and protein-coding regions was calculated using DnaSP v6.12.03 (Rozas et al., 2017). Sliding-window analyses were conducted with a window size of 600 bp and step size of 200 bp for individual genes, while whole-genome variability was assessed using default batch settings.
For phylogenetic reconstruction, the newly sequenced chloroplast genomes of T. biflora and T. turkestanica were aligned with available Tulipa and related genera chloroplast genomes retrieved from GenBank. Maximum likelihood (ML) phylogenetic analysis was performed using RAxML v8.2.12 (Stamatakis, 2014) with the GTR + I + G4 substitution model. Branch support was assessed using 1,000 bootstrap replicates.
ResultsРезультатыНәтижелер
Chloroplast Genome Organization and Gene Content
The complete chloroplast genomes of T. turkestanica and T. biflora were 152,025 bp and 152,022 bp in length, respectively. Both genomes exhibited the conserved quadripartite structure characteristic of angiosperms, comprising a large single-copy (LSC) region, a small single-copy (SSC) region, and two inverted repeat (IR) regions (Fig. 1). Each genome encoded 84 protein-coding genes with identical gene composition (Appendix 1). The high degree of structural conservation reflects the overall stability of plastome architecture within Tulipa.
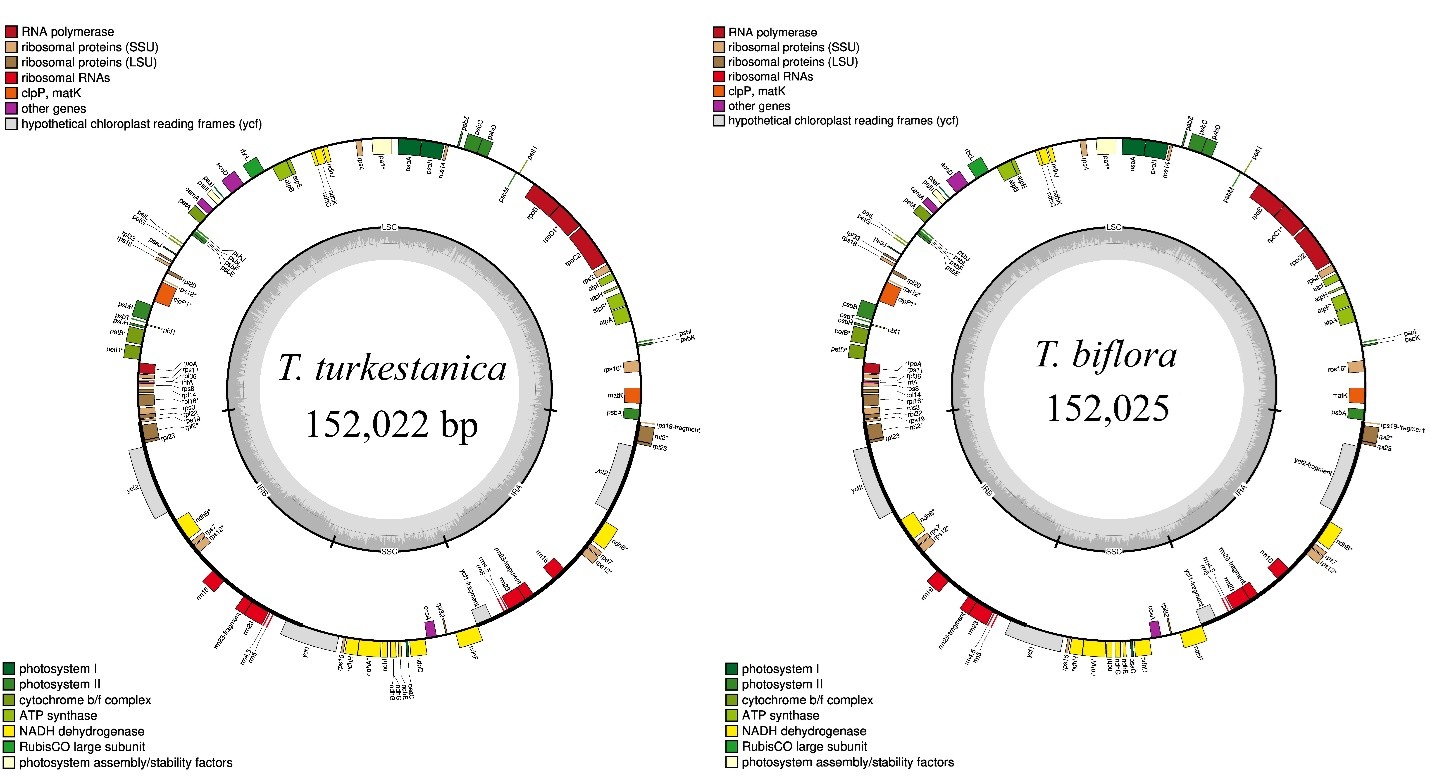
Chloroplast genome maps of T. turkestanica and T. biflora. Genes shown on the outside of the circle are transcribed clockwise, while those on the inside are transcribed counterclockwise. Genes are color-coded by functional category. Chloroplast genome maps of T. turkestanica and T. biflora. Genes shown on the outside of the circle are transcribed clockwise, while those on the inside are transcribed counterclockwise. Genes are color-coded by functional category. Chloroplast genome maps of T. turkestanica and T. biflora. Genes shown on the outside of the circle are transcribed clockwise, while those on the inside are transcribed counterclockwise. Genes are color-coded by functional category.
Nucleotide Diversity
Nucleotide diversity analysis revealed that the petD-rpoA intergenic spacer region exhibited the highest variability (π = 0.01625), suggesting this region as a potential molecular marker for population genetic studies (Fig. 2). Protein-coding regions showed generally lower diversity, consistent with functional constraints on these sequences.
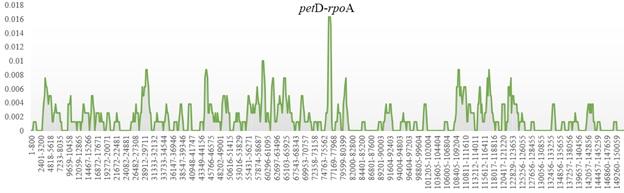
Nucleotide diversity (π) analysis of T. turkestanica and T. biflora chloroplast genomes. The petD-rpoA region shows the highest variability. Nucleotide diversity (π) analysis of T. turkestanica and T. biflora chloroplast genomes. The petD-rpoA region shows the highest variability. Nucleotide diversity (π) analysis of T. turkestanica and T. biflora chloroplast genomes. The petD-rpoA region shows the highest variability.
Simple Sequence Repeat Analysis
A total of 88 SSR motifs representing 23 distinct types were identified in T. turkestanica, whereas 71 SSR motifs representing 17 types were detected in T. biflora (Fig. 3). In both species, mononucleotide repeats were the most abundant, with adenine/thymine (A/T) motifs predominating. Specifically, A motifs occurred 29 times in T. turkestanica and 31 times in T. biflora, while T motifs were found 23 times in T. turkestanica and 24 times in T. biflora.
Dinucleotide repeats showed marked differences between the two species. In T. turkestanica, the AT motif was most common (10 occurrences), whereas only one AT repeat was observed in T. biflora. Furthermore, other dinucleotide motifs (AG, CT, GA, TA, and TC) were present at low frequencies in T. turkestanica but were absent or rare in T. biflora. AG/CT repeats, in particular, were unique to T. turkestanica with seven occurrences, representing a species-specific marker.
Trinucleotide and higher-order repeats (tetranucleotide and pentanucleotide) were detected in both species but at comparatively low frequencies. Common motifs included ATA, TAT, TTA, AATA, AGAA, ATAA, ATCA, ATTT, GAAT, TAAT, TAGA, and TATT. Both plastomes exhibited a strong bias toward A/T-rich repeats, consistent with the high A/T content of chloroplast genomes. However, the greater number and diversity of SSRs in T. turkestanica suggests differential microsatellite accumulation dynamics between the two species.
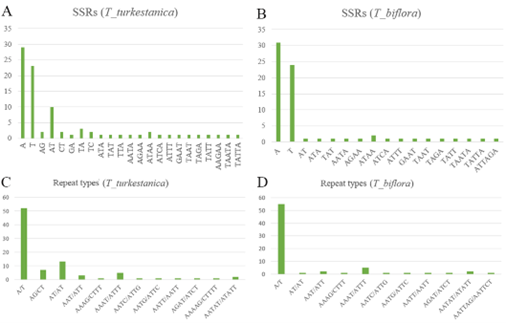
Simple sequence repeat (SSR) analysis of T. turkestanica and T. biflora. (A) SSR motifs in T. turkestanica, (B) SSR motifs in T. biflora, (C) SSR repeat types in T. turkestanica, and (D) SSR repeat types in T. biflora. Simple sequence repeat (SSR) analysis of T. turkestanica and T. biflora. (A) SSR motifs in T. turkestanica, (B) SSR motifs in T. biflora, (C) SSR repeat types in T. turkestanica, and (D) SSR repeat types in T. biflora. Simple sequence repeat (SSR) analysis of T. turkestanica and T. biflora. (A) SSR motifs in T. turkestanica, (B) SSR motifs in T. biflora, (C) SSR repeat types in T. turkestanica, and (D) SSR repeat types in T. biflora.
Phylogenetic Relationships
Maximum likelihood phylogenetic analysis based on complete chloroplast genomes resolved relationships within subgenus Eriostemones with strong bootstrap support (Fig. 4). Tulipa biflora formed a well-supported clade with T. sogdiana (BS = 100), while T. turkestanica clustered with two accessions of T. buhseana, also with maximum support (BS = 100). These results indicate close evolutionary relationships among these taxa and confirm their placement within distinct lineages of subgenus Eriostemones.
The phylogenetic topology demonstrated clear separation of additional species within the subgenus, while T. sinkiangensis (subgenus Orithyia) was clearly distinguished as the outgroup. Overall, the chloroplast genome-based phylogeny provides robust resolution of relationships within Eriostemones and supports the systematic placement of T. biflora and T. turkestanica within well-defined evolutionary lineages.

Maximum likelihood phylogenetic tree based on complete chloroplast genomes. T. sinkiangensis from subgenus Orithyia was used as the outgroup. Bootstrap support values (≥50%) are shown at nodes. Maximum likelihood phylogenetic tree based on complete chloroplast genomes. T. sinkiangensis from subgenus Orithyia was used as the outgroup. Bootstrap support values (≥50%) are shown at nodes. Maximum likelihood phylogenetic tree based on complete chloroplast genomes. T. sinkiangensis from subgenus Orithyia was used as the outgroup. Bootstrap support values (≥50%) are shown at nodes.
DiscussionОбсуждениеТалқылау
Chloroplast Genome Conservation in Tulipa
The complete chloroplast genomes of T. turkestanica and T. biflora exhibited highly conserved structural organization, including the typical quadripartite architecture (LSC, SSC, and two IR regions) and nearly identical genome sizes. The presence of 84 protein-coding genes in both species, with complete gene content conservation, reflects the structural stability of chloroplast genomes across the genus Tulipa. This pattern is consistent with previous comparative studies in Liliaceae, which have documented limited genome rearrangements and gene losses within the family (She et al., 2020). Such structural uniformity indicates that major genomic reorganization events have played a minimal role in plastome evolution within this lineage.
Microsatellite Variation and Species Differentiation
Despite overall structural conservation, notable variation was observed in microsatellite composition between the two species. Both plastomes were strongly enriched in A/T-rich SSRs, a pattern widely documented in plant chloroplast genomes and generally attributed to replication slippage and the inherent AT bias of plastid DNA (Oliveira et al., 2006). However, T. turkestanica exhibited a higher total number (88 vs. 71) and greater diversity (23 vs. 17 types) of SSR motifs compared to T. biflora, suggesting differential accumulation of microsatellites during evolutionary divergence.
The most striking difference was observed in dinucleotide repeats. T. turkestanica displayed significantly higher frequencies of AT motifs (10 occurrences) and possessed unique AG/CT repeats (7 occurrences), which were absent in T. biflora. These species-specific differences in repeat composition indicate distinct mutation dynamics and may reflect different evolutionary histories or population genetic processes. Such SSR polymorphisms represent valuable molecular markers that could be developed for population genetic studies, phylogeographic analyses, and species authentication within Tulipa (Raskina, 2017). The petD-rpoA intergenic region, showing the highest nucleotide diversity (π = 0.01625), provides an additional target for developing informative markers for evolutionary and conservation studies.
Phylogenetic Insights and Evolutionary Implications
Phylogenetic reconstruction based on complete chloroplast genomes provided robust resolution within subgenus Eriostemones. The close clustering of T. biflora with T. sogdiana and of T. turkestanica with T. buhseana corroborates morphological affinities previously proposed for these taxa (Dekhkonov et al., 2022) and confirms their placement within distinct evolutionary lineages. However, this chloroplast genome-based topology contrasts with previous phylogenies based on nuclear ITS markers (Kubentayev et al., 2024), suggesting the influence of introgression and incomplete lineage sorting.
Cytonuclear discordance is a common phenomenon in Liliaceae and has been extensively documented in recent phylogenomic studies of tribe Tulipeae (Zhang et al., 2025). This discordance typically arises from processes such as ancient hybridization events, asymmetric gene flow, and incomplete sorting of ancestral polymorphisms. In the case of Central Asian tulips, the complex geological history of the region, including repeated climatic oscillations and geographic fragmentation, likely promoted multiple opportunities for interspecific gene flow while simultaneously facilitating lineage diversification. The clear separation of T. sinkiangensis (subgenus Orithyia) from subgenus Eriostemones in our phylogeny supports the major subgeneric divisions within Tulipa and validates the use of chloroplast genomes for resolving higher-level systematic relationships.
Conservation and Future Directions
The comparative chloroplast genome analysis of T. biflora and T. turkestanica demonstrates that while plastome structure and gene content are highly conserved, repeat sequences and intergenic spacer regions provide informative variation for species differentiation and evolutionary studies. Given the ongoing taxonomic challenges and conservation concerns surrounding wild tulips, with 62 species currently listed on the IUCN Red List, the genomic resources generated in this study represent a valuable foundation for future research. These resources can support species delimitation efforts, facilitate the development of molecular markers for population genetic monitoring, enable phylogeographic studies to identify evolutionarily significant units, and inform evidence-based conservation strategies for these threatened species. Further integration of nuclear genome data with chloroplast genomic information will be essential for fully resolving the complex evolutionary history and systematic relationships within this ecologically and culturally important genus.
Chloroplast Genome Conservation in Tulipa
The complete chloroplast genomes of T. turkestanica and T. biflora exhibited highly conserved structural organization, including the typical quadripartite architecture (LSC, SSC, and two IR regions) and nearly identical genome sizes. The presence of 84 protein-coding genes in both species, with complete gene content conservation, reflects the structural stability of chloroplast genomes across the genus Tulipa. This pattern is consistent with previous comparative studies in Liliaceae, which have documented limited genome rearrangements and gene losses within the family (She et al., 2020). Such structural uniformity indicates that major genomic reorganization events have played a minimal role in plastome evolution within this lineage.
Microsatellite Variation and Species Differentiation
Despite overall structural conservation, notable variation was observed in microsatellite composition between the two species. Both plastomes were strongly enriched in A/T-rich SSRs, a pattern widely documented in plant chloroplast genomes and generally attributed to replication slippage and the inherent AT bias of plastid DNA (Oliveira et al., 2006). However, T. turkestanica exhibited a higher total number (88 vs. 71) and greater diversity (23 vs. 17 types) of SSR motifs compared to T. biflora, suggesting differential accumulation of microsatellites during evolutionary divergence.
The most striking difference was observed in dinucleotide repeats. T. turkestanica displayed significantly higher frequencies of AT motifs (10 occurrences) and possessed unique AG/CT repeats (7 occurrences), which were absent in T. biflora. These species-specific differences in repeat composition indicate distinct mutation dynamics and may reflect different evolutionary histories or population genetic processes. Such SSR polymorphisms represent valuable molecular markers that could be developed for population genetic studies, phylogeographic analyses, and species authentication within Tulipa (Raskina, 2017). The petD-rpoA intergenic region, showing the highest nucleotide diversity (π = 0.01625), provides an additional target for developing informative markers for evolutionary and conservation studies.
Phylogenetic Insights and Evolutionary Implications
Phylogenetic reconstruction based on complete chloroplast genomes provided robust resolution within subgenus Eriostemones. The close clustering of T. biflora with T. sogdiana and of T. turkestanica with T. buhseana corroborates morphological affinities previously proposed for these taxa (Dekhkonov et al., 2022) and confirms their placement within distinct evolutionary lineages. However, this chloroplast genome-based topology contrasts with previous phylogenies based on nuclear ITS markers (Kubentayev et al., 2024), suggesting the influence of introgression and incomplete lineage sorting.
Cytonuclear discordance is a common phenomenon in Liliaceae and has been extensively documented in recent phylogenomic studies of tribe Tulipeae (Zhang et al., 2025). This discordance typically arises from processes such as ancient hybridization events, asymmetric gene flow, and incomplete sorting of ancestral polymorphisms. In the case of Central Asian tulips, the complex geological history of the region, including repeated climatic oscillations and geographic fragmentation, likely promoted multiple opportunities for interspecific gene flow while simultaneously facilitating lineage diversification. The clear separation of T. sinkiangensis (subgenus Orithyia) from subgenus Eriostemones in our phylogeny supports the major subgeneric divisions within Tulipa and validates the use of chloroplast genomes for resolving higher-level systematic relationships.
Conservation and Future Directions
The comparative chloroplast genome analysis of T. biflora and T. turkestanica demonstrates that while plastome structure and gene content are highly conserved, repeat sequences and intergenic spacer regions provide informative variation for species differentiation and evolutionary studies. Given the ongoing taxonomic challenges and conservation concerns surrounding wild tulips, with 62 species currently listed on the IUCN Red List, the genomic resources generated in this study represent a valuable foundation for future research. These resources can support species delimitation efforts, facilitate the development of molecular markers for population genetic monitoring, enable phylogeographic studies to identify evolutionarily significant units, and inform evidence-based conservation strategies for these threatened species. Further integration of nuclear genome data with chloroplast genomic information will be essential for fully resolving the complex evolutionary history and systematic relationships within this ecologically and culturally important genus.
Chloroplast Genome Conservation in Tulipa
The complete chloroplast genomes of T. turkestanica and T. biflora exhibited highly conserved structural organization, including the typical quadripartite architecture (LSC, SSC, and two IR regions) and nearly identical genome sizes. The presence of 84 protein-coding genes in both species, with complete gene content conservation, reflects the structural stability of chloroplast genomes across the genus Tulipa. This pattern is consistent with previous comparative studies in Liliaceae, which have documented limited genome rearrangements and gene losses within the family (She et al., 2020). Such structural uniformity indicates that major genomic reorganization events have played a minimal role in plastome evolution within this lineage.
Microsatellite Variation and Species Differentiation
Despite overall structural conservation, notable variation was observed in microsatellite composition between the two species. Both plastomes were strongly enriched in A/T-rich SSRs, a pattern widely documented in plant chloroplast genomes and generally attributed to replication slippage and the inherent AT bias of plastid DNA (Oliveira et al., 2006). However, T. turkestanica exhibited a higher total number (88 vs. 71) and greater diversity (23 vs. 17 types) of SSR motifs compared to T. biflora, suggesting differential accumulation of microsatellites during evolutionary divergence.
The most striking difference was observed in dinucleotide repeats. T. turkestanica displayed significantly higher frequencies of AT motifs (10 occurrences) and possessed unique AG/CT repeats (7 occurrences), which were absent in T. biflora. These species-specific differences in repeat composition indicate distinct mutation dynamics and may reflect different evolutionary histories or population genetic processes. Such SSR polymorphisms represent valuable molecular markers that could be developed for population genetic studies, phylogeographic analyses, and species authentication within Tulipa (Raskina, 2017). The petD-rpoA intergenic region, showing the highest nucleotide diversity (π = 0.01625), provides an additional target for developing informative markers for evolutionary and conservation studies.
Phylogenetic Insights and Evolutionary Implications
Phylogenetic reconstruction based on complete chloroplast genomes provided robust resolution within subgenus Eriostemones. The close clustering of T. biflora with T. sogdiana and of T. turkestanica with T. buhseana corroborates morphological affinities previously proposed for these taxa (Dekhkonov et al., 2022) and confirms their placement within distinct evolutionary lineages. However, this chloroplast genome-based topology contrasts with previous phylogenies based on nuclear ITS markers (Kubentayev et al., 2024), suggesting the influence of introgression and incomplete lineage sorting.
Cytonuclear discordance is a common phenomenon in Liliaceae and has been extensively documented in recent phylogenomic studies of tribe Tulipeae (Zhang et al., 2025). This discordance typically arises from processes such as ancient hybridization events, asymmetric gene flow, and incomplete sorting of ancestral polymorphisms. In the case of Central Asian tulips, the complex geological history of the region, including repeated climatic oscillations and geographic fragmentation, likely promoted multiple opportunities for interspecific gene flow while simultaneously facilitating lineage diversification. The clear separation of T. sinkiangensis (subgenus Orithyia) from subgenus Eriostemones in our phylogeny supports the major subgeneric divisions within Tulipa and validates the use of chloroplast genomes for resolving higher-level systematic relationships.
Conservation and Future Directions
The comparative chloroplast genome analysis of T. biflora and T. turkestanica demonstrates that while plastome structure and gene content are highly conserved, repeat sequences and intergenic spacer regions provide informative variation for species differentiation and evolutionary studies. Given the ongoing taxonomic challenges and conservation concerns surrounding wild tulips, with 62 species currently listed on the IUCN Red List, the genomic resources generated in this study represent a valuable foundation for future research. These resources can support species delimitation efforts, facilitate the development of molecular markers for population genetic monitoring, enable phylogeographic studies to identify evolutionarily significant units, and inform evidence-based conservation strategies for these threatened species. Further integration of nuclear genome data with chloroplast genomic information will be essential for fully resolving the complex evolutionary history and systematic relationships within this ecologically and culturally important genus.
AcknowledgmentsБлагодарностиАлғыс сөздер
This research was supported by [funding information to be added]. We thank the staff of the TASH herbarium for assistance with specimen deposition and the field workers who helped with sample collection.
Data & Code AvailabilityДоступность данных и кодаДеректер мен кодқа қолжетімділік
- Data: The complete chloroplast genome sequences have been deposited in GenBank under accession numbers [to be added].
- Code: Analysis scripts are available upon request from the corresponding author.
EthicsЭтикаЭтика
Sample collection was conducted in accordance with local regulations. Voucher specimens were deposited in TASH herbarium.
Competing InterestsКонфликт интересовМүдделер қақтығысы
The authors declare no competing interests.
References
- Asatulloev, T., Dekhkonov, D., Yusupov, Z., Tojiboeva, U., Cai, L., Tojibaev, K., Sun, W., 2023. Ecoregional and phytogeographical insights into the distribution of Tulipa in the 'Nature Imperiled' area of Central Asia for effective conservation. Diversity 15, 1195. DOI: 10.3390/d15121195
- Beier, S., Thiel, T., Münch, T., Scholz, U., Mascher, M., 2017. MISA-web: a web server for microsatellite prediction. Bioinformatics 33, 2583–2585. DOI: 10.1093/bioinformatics/btx198
- Botschantzeva, Z., 1962. Tulips: taxonomy, morphology, cytology, phytogeography and physiology. Russian edition; English translation by Varekamp, H. (1982). Rotterdam: Balkema.
- Christenhusz, M.J.M., Govaerts, R., David, J.C., Hall, T., Borland, K., Roberts, P.S., Tuomisto, A., Buerki, S., Chase, M.W., Fay, M.F., 2013. Tiptoe through the tulips – cultural history, molecular phylogenetics and classification of Tulipa (Liliaceae). Botanical Journal of the Linnean Society 172, 280–328. DOI: 10.1111/boj.12061
- de Groot, J.J., Zonneveld, B.J.M., 2024. Three new species of Tulipa from Central Asia. International Rock Gardener 168, 3–18.
- Dekhkonov, D., Tojibaev, K., Yusupov, Z., Makhmudjanov, D., Asatulloev, T., 2022. Morphology of tulips (Tulipa, Liliaceae) in its primary centre of diversity. Plant Diversity of Central Asia 1, 52–70. DOI: 10.54981/PDCA/vol1_iss1/a3
- Hall, A.D., 1940. The genus Tulipa. London: The Royal Horticultural Society.
- International Union for Conservation of Nature (IUCN), 2022. The IUCN Red List of Threatened Species, Version 2021–3. https://www.iucnredlist.org
- Jin, J.J., Yu, W.B., Yang, J.B., Song, Y., DePamphilis, C.W., Yi, T.S., Li, D.Z., 2020. GetOrganelle: a fast and versatile toolkit for accurate de novo assembly of organelle genomes. Genome Biology 21, 241. DOI: 10.1186/s13059-020-02154-5
- Kearse, M., Moir, R., Wilson, A., Stones-Havas, S., Cheung, M., Sturrock, S., Buxton, S., Cooper, A., Markowitz, S., Duran, C., Thierer, T., Ashton, B., Meintjes, P., Drummond, A., 2012. Geneious Basic: an integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics 28, 1647–1649. DOI: 10.1093/bioinformatics/bts199
- Kubentayev, S., Yermagambetova, M., Vesselova, P., Turuspekov, Y., Abugalieva, S., 2024. Phylogenetic relationships among Tulipa species from Kazakhstan based on ITS sequences. Plant Systematics and Evolution 310, 15. DOI: 10.1007/s00606-024-01889-0
- Lohse, M., Drechsel, O., Bock, R., 2007. OrganellarGenomeDRAW (OGDRAW): a tool for the easy generation of high-quality custom graphical maps of plastid and mitochondrial genomes. Current Genetics 52, 267–274. DOI: 10.1007/s00294-007-0161-y
- Miao, Y., Herrmann, M., Wu, F., Yan, X., Yang, S., 2012. What controlled mid–late Miocene long-term aridification in Central Asia? Global cooling or Tibetan Plateau uplift: a review. Earth-Science Reviews 112, 155–172. DOI: 10.1016/j.earscirev.2012.02.003
- Myers, N., Mittermeier, R.A., Mittermeier, C.G., da Fonseca, G.A.B., Kent, J., 2000. Biodiversity hotspots for conservation priorities. Nature 403, 853–858. DOI: 10.1038/35002501
- Oliveira, E.J., Pádua, J.G., Zucchi, M.I., Vencovsky, R., Vieira, M.L.C., 2006. Origin, evolution and genome distribution of microsatellites. Genetics and Molecular Biology 29, 294–307. DOI: 10.1590/S1415-47572006000200018
- POWO, 2025. Plants of the World Online. Royal Botanic Gardens, Kew. https://powo.science.kew.org (accessed 07 August 2025).
- Raskina, O., 2017. Genotype- and cell-specific dynamics of tandem repeat patterns in Aegilops speltoides (Poaceae, Triticeae). Cytogenetic and Genome Research 153, 105–116. DOI: 10.1159/000484562
- Rozas, J., Ferrer-Mata, A., Sánchez-DelBarrio, J.C., Guirao-Rico, S., Librado, P., Ramos-Onsins, S.E., Sánchez-Gracia, A., 2017. DnaSP 6: DNA sequence polymorphism analysis of large data sets. Molecular Biology and Evolution 34, 3299–3302. DOI: 10.1093/molbev/msx248
- She, R., Zhao, P., Zhou, H., Yue, M., Yan, F., Hu, G., Zhang, S., et al., 2020. Complete chloroplast genomes of Liliaceae (s.l.) species: comparative genomic and phylogenetic analyses. Nordic Journal of Botany 38, e02545. DOI: 10.1111/njb.02545
- Stamatakis, A., 2014. RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30, 1312–1313. DOI: 10.1093/bioinformatics/btu033
- Vvedensky, A.I., Kovalevskaya, S.S., 1971. Tulipa L. In: Vvedensky, A.I. (Ed.), Conspectus Florae Asiae Mediae, Vol. 2. Tashkent: Academy of Sciences of UzSSR, pp. 94–109.
- Zhang, Z., Wang, M., Yang, Z., Comes, H.P., Zhong, X., Folk, R.A., Li, P., et al., 2025. Incomplete lineage sorting and introgression among genera and species of Liliaceae tribe Tulipeae: insights from phylogenomics. BMC Biology 23, 1–22. DOI: 10.1186/s12915-025-02204-z
- Zonneveld, B.J.M., 2009. The systematic value of nuclear genome size for all species of Tulipa L. (Liliaceae). Plant Systematics and Evolution 281, 217–245. DOI: 10.1007/s00606-009-0203-7
Appendix 1
| Category | Gene Group | Gene Name | Number |
|---|---|---|---|
| Photosynthesis | Photosystem I subunits | psaB, psaA, psaI, psaJ, psaC | 5 |
| Photosystem II subunits | psbA, psbK, psbI, psbM, psbD, psbC, psbZ, psbJ, psbL, psbF, psbE, psbB, psbT, psbH | 14 | |
| NADH dehydrogenase subunits | ndhJ, ndhK, ndhC, ndhB (2)#, ndhH, ndhA#, ndhI, ndhG (2), ndhE, ndhD, ndhF | 13 | |
| Cytochrome b/f complex subunits | petN, petA, petL, petG, petB#, petD# | 6 | |
| RuBisCO large subunit | rbcL | 1 | |
| ATP synthase subunits | atpA, atpF#, atpH, atpI, atpE, atpB | 6 | |
| Self-replication | Large ribosomal subunit proteins | rpl33, rpl20, rpl36, rpl14, rpl16#, rpl22, rpl2 (2)#, rpl23 (2), rpl32 | 11 |
| Small ribosomal subunit proteins | rps12 (2)##, rps16#, rps2, rps14, rps4, rps18, rps11, rps8, rps3, rps19, rps7 (2), rps15, rps19-fragment | 15 | |
| RNA polymerase subunits | rpoC2, rpoC1#, rpoB, rpoA | 4 | |
| Ribosomal RNAs | rrn16 (2), rrn23-fragment (2), rrn23 (2), rrn4.5 (2), rrn5 (2) | 10 | |
| Other genes | Maturase | matK | 1 |
| Protease | clpP1## | 1 | |
| Envelope membrane protein | cemA | 1 | |
| Acetyl-CoA carboxylase | accD | 1 | |
| Cytochrome c synthesis | ccsA | 1 | |
| Translation initiation factor | infA | 1 | |
| Conserved open reading frames | ycf2 (2), ycf1, ycf1-fragment | 4 | |
| Other genes | pafI##, pafII, pbf1 | 3 |